Moran Benhar, Ph.D.
 Department of Biochemistry | Faculty of Medicine
Department of Biochemistry | Faculty of Medicine
Rappaport Building, 6th floor
Tel: +972-4-829-5376
Fax: +972-4-829-5412
E-mail: benhar@technion.ac.il
פרופסור מורן בנהר – מחלקה לביוכימיה
⇒ Open positions are available for Master and PhD students
למתעניינים במחקר במסגרת לימודי מסטר ודוקטורט – צרו קשר
See more information here
The main goal of our lab is to discover how oxidant and antioxidant systems regulate key signaling pathways in cancer cells and macrophages. We focus on understanding how redox modifications of proteins influence inflammatory and cell death pathways.
Mammalian cells continuously generate oxidants as part of normal aerobic metabolism and in response to diverse external stimuli (growth factors, hormones, stress signals, etc.). Several types of biological ‘reactive species’ are known and named according to the nature of the reactive atom, that is, oxygen, nitrogen or sulfur: reactive oxygen species (ROS), reactive nitrogen species (RNS) and reactive sulfur species (RSS). Notable examples include superoxide (O2⋅–), hydrogen peroxide (H2O2), nitric oxide (NO) and hydrogen sulfide (H2S).⋅ The major antioxidant systems, the glutathione and thioredoxin systems, play key roles in protecting cells from oxidative stress and regulation of ROS signaling.
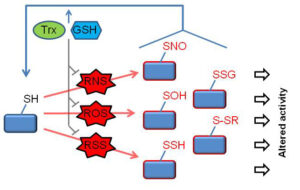
Cysteines confer redox regulation of protein function, namely the reversible post-translational modification that alters protein activity as a result of change in its oxidation state. In response to changes in cellular levels of ROS, RNS or RSS cysteine thiol (-SH) groups undergo a spectrum of covalent modifications (see Figure). These reversible thiol modifications are increasingly recognized to regulate a wide range of cellular functions, such as proliferation, migration, differentiation, and death.
Functional modifications of the thiol proteome. In response to generation of reactive oxygen/nitrogen/sulfur species (ROS/RNS/RSS), protein cysteine thiols (R–SH) undergo a range of oxidative modifications. Of these, major reversible modifications include nitrosylation (R–SNO) (induced by RNS), sulfenylation (R–SOH) (induced by ROS) and persulfidation (R–SSH) (induced by RSS). Glutathione (GSH), thioredoxin (Trx) and associated redox enzymes govern the redox state of proteins by two mechanisms, (1) by eliminating ROS/RNS/RSS), and (2) by catalyzing the reduction of oxidized/nitrosylated/persulfidated thiols. Accordingly, the balance between oxidants and GSH/Trx determines the magnitude and duration of each of the redox modifications and thus the downstream functional consequences.
We are particularly interested in redox regulation through S-nitrosylation, the attachment of NO moiety to cysteine thiol. Our research team is studying the role of S-nitrosylation in cellular signaling, inflammation and cancer. We employ global (proteomics) as well as more directed and targeted approaches to discover novel nitrosylation-based control mechanisms.
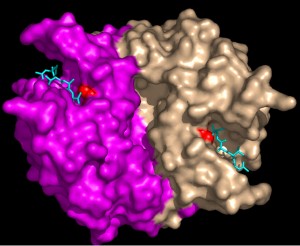
Structural model of S-nitrosylated caspase-3. Binding of the nitric oxide group (red color) to the active site cysteine inhibits caspase activity resulting in protection from programmed cell death (apoptosis).
Among the gaseous signaling molecules, similar to NO, hydrogen sulfide (H2S) as recently emerged has a pleiotropic signaling molecule the affects diverse cellular processes. Cellular effects of H2S are mediated in part via thiol persulfidation (Cys-SSH), also known as ‘sulfhydration’.
Our research goals are:
1. Identify novel targets of NO and H2S in macrophages and cancer cells
2. Elucidate how H2S regulates cancer cell death
3. Develop new proteomic methods to analyze protein oxidation.
4. Characterization of the thiol redox proteome in the macrophage inflammatory response.
5. Elucidate the role of antioxidant systems in regulating protein nitrosylation and persulfidation